Protein Folding Imaging Services
Protein is a complex organic compound composed of amino acids. Due to the hydrophilicity, hydrophobicity, positive charge, negative charge and other characteristics of amino acid residues, it can be folded into a precise 3D structure through the interaction between the residues. The sequence of amino acids determines the unique 3D structure of each protein and its specific functions. Protein participates in almost every process in the cell and is necessary to maintain the structure and function of human tissues and organs. Protein folding errors can induce a variety of degenerative diseases, such as Parkinson's disease, Alzheimer's disease, frostbite, and senile heart failure. Therefore, imaging analysis of the folding process of proteins is of great significance for the early diagnosis and treatment of diseases.
 Figure 1. Protein folding on the ribosome (Balchin, et al., 2016).
Figure 1. Protein folding on the ribosome (Balchin, et al., 2016).
Protein Folding Imaging Analysis
Most proteins must be folded into a unique three-dimensional structure to perform their biological functions. However, in the cellular environment, newly synthesized proteins may misfold and form toxic aggregates. To ensure effective folding, it is necessary to monitor the dynamic process of protein folding in real time. Fluorescence imaging technology provides a useful and reliable method for elucidating the mechanism of protein folding and aggregation.
CD BioSciences has been committed to the research and development of fluorescent probes and imaging technology for many years. We can provide you with folded protein covalent probes to visually analyze protein dimerization, interaction and conformational changes. Understanding the dynamic process of protein folding and aggregation can help you diagnose and treat diseases caused by protein folding errors.
Protein Folding Imaging Analysis Workflow
The following are the specific steps of protein folding imaging analysis:

Synthesis of fluorescent probe
Step 1
Fluorescent label
Step 2
Imaging
Step 3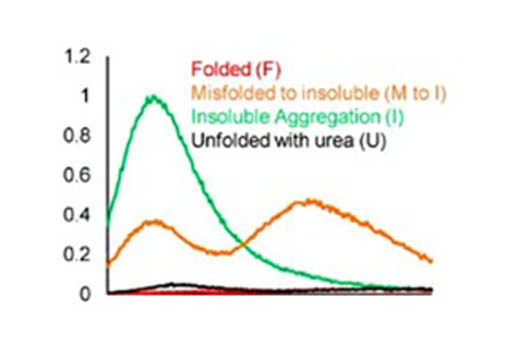
Image analysis
Step 4Delivery
Fluorescence spectra of fluorescent probes in folded, misfolded and aggregated proteins
Fluorescence quantitative thermal displacement curve
Relative fluorescence intensity
Total number of cell aggregates
Percentage of cells with aggregates
Aggregates of each cell
Aggregate area and location aggregate (cytoplasmic, perinuclear or nuclear)
Our Advantages
- High signal-to-noise ratio, high resolution
- Able to perform real-time dynamic imaging of protein folding
- Experienced scientists provide experimental consultation
- Reasonable price and short turnaround time
CD BioSciences has a professional team and advanced imaging equipment. The entire process of protein folding imaging analysis is operated by experienced technicians to ensure the accuracy of the experiment. If you have any needs, please feel free to contact us.
- Klickstein, Jacob Aaron, Sirisha Mukkavalli, and Malavika Raman. "AggreCount: an unbiased image analysis tool for identifying and quantifying cellular aggregates in a spatially defined manner." Journal of Biological Chemistry 295.51 (2020): 17672-17683.
- Balchin, David, Manajit Hayer-Hartl, and F. Ulrich Hartl. "In vivo aspects of protein folding and quality control." Science 353.6294 (2016).
- Wan, Wang, et al. "Covalent Probes for Aggregated Protein Imaging via Michael Addition." Angewandte Chemie International Edition 60.20 (2021): 11335-11343.
*If your organization requires the signing of a confidentiality agreement, please contact us by email.
Please note: Our services can only be used for research purposes. Do not use in diagnostic or therapeutic procedures!

